Caracterización de bacterias degradadoras de hidrocarburos en la estación EPEA, costa del Atlántico Sur
DOI:
https://doi.org/10.47193/mafis.3412021010304Palabras clave:
PAH, biorremediación, biosurfactanteResumen
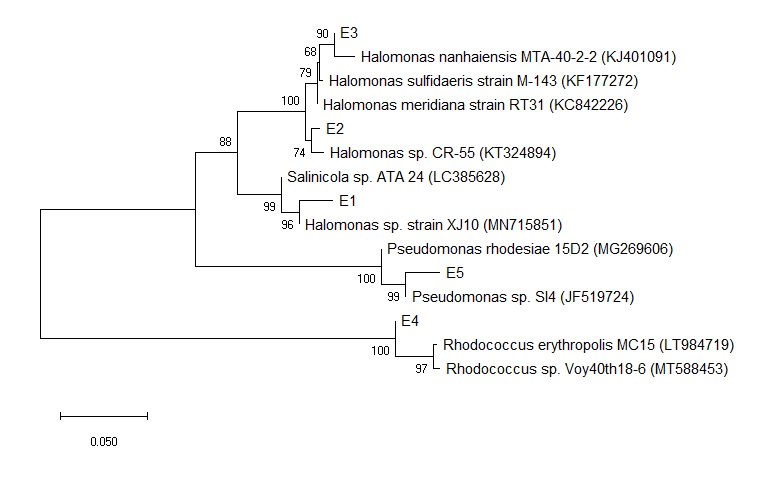
Las bacterias degradadoras de hidrocarburos (BDH) fueron monitoreadas desde 2006 a 2018 en la Estación Permanente de Estudios Ambientales (EPEA), con el fin de analizar su abundancia y la potencialidad de metabolizar estos contaminantes. La presencia de BDH se detectó con valores de recuento que oscilaron entre 103 y 105 UFC ml-1. Se observó un ligero aumento a lo largo del tiempo, que podría estar relacionado con cambios en la temperatura marina reportados en los últimos años. Se analizaron 36 BDH para determinar su crecimiento en varios hidrocarburos y algunas de ellas mostraron un perfil de biodegradación amplio. Además, a partir de cultivos de enriquecimiento con fenantreno (Phe), se identificaron filogenéticamente cinco cepas como Halomonas sp. E1, E2 y E3; Rhodococcus sp. E4 y Pseudomonas sp. E5. Se demostró una degradación completa de Phe para las cepas E4 y E5, mientras que las cepas E1, E2, E3 y E4 mostraron producción de surfactante. Este estudio contribuyó con el primer conocimiento sobre el potencial intrínseco de biodegradación de los hidrocarburos por las comunidades bacterianas en EPEA. Algunas de las cepas exhibieron propiedades fisiológicas que pueden tener importancia ecológica sobre alteraciones ambientales como la presencia de contaminantes. En particular, Rhodococcus sp. E4 podría ser una alternativa para la selección microbiana en la degradación de hidrocarburos poliaromáticos. Se necesitan más estudios para evaluar el impacto del cambio climático en los procesos de desintoxicación mediados por microbios.
Descargas
Referencias
Abo-State MAM, El-Dars FM, Abdin BA. 2018. Isolation and identification of pyrene degrading bacteria and its pathway from Suez Oil Processing Company, Suez, Egypt. J Eco Heal Env. 6: 63-76.
Ali I, Prasongsuk S, Akbar A, Aslam M, Lotrakul P, Punnapayak H, Rakshit SK. 2016. Hypersaline habitats and halophilic microorganisms. Maejo Int J Sci Technol. 10: 330-345.
Batista SB, Mounteer AH, Amorim FR, Totola MR. 2006. Isolation and characterization of biosurfactant/bioemulsifier-producing bacteria from petroleum contaminated sites. Bioresour Technol. 97: 868-875.
Bogardt AH, Hemmingsen BB. 1992. Enumeration of phenanthrene-degrading bacteria by an overlayer technique and its use in evaluation of petroleum-contaminated sites. Appl Environ Microbiol. 58: 2579-2582.
Brzeszcz J, Kaszycki P. 2018. Aerobic bacteria degrading both n-alkanes and aromatic hydrocarbons: an undervalued strategy for metabolic diversity and flexibility. Biodegradation. 29: 359-407.
Cabral H, Fonseca V, Sousa T, Costa Leal M. 2019. Synergistic effects of climate change and marine pollution: an overlooked interaction in coastal and estuarine areas. Int J Environ Res. 16: 2737.
Cappello S, Caruso G, Zampino D, Monticelli LS, Maimone G, Denaro R, Tripodo B, Troussellier M, Yakimov MM, Giuliano L. 2007. Microbial community dynamics during assays of harbour oil spill bioremediation: a microscale simulation study. J Appl Microbiol. 102: 184-194.
Coelho FJ, Cleary DFR, Costa R, Ferreira M, Polonia ARM, Silva AMS, Simoes MMQ, Oliveira V, Gomes NCM. 2016. Multitaxon activity profiling reveals differential microbial response to reduced seawater pH and oil pollution. Mol Ecol. 25: 4645-4659.
Coelho FJ, Santos AL, Coimbra J, Almeida A, Cunha A, Cleary DFR, Calado R, Gomes NCM. 2013. Interactive effects of global climate change and pollution on marine microbes: the way ahead. Ecol Evol. 3: 1808-1818.
Corti Monzón G, Nisenbaum M, Herrera Seitz MK, Murialdo SE. 2018. New findings on aromatic compounds’ degradation and their metabolic pathways, the biosurfactant production and motility of the halophilic bacterium Halomonas sp. KHS3. Curr. Microbiol. 75: 1108-1118.
Cui Z, Lai Q, Dong CH, Shao Z. 2008. Biodiversity of polycyclic aromatic hydrocarbon-degrading bacteria from deep sea sediments of the Middle Atlantic Ridge. Environ Microbiol. 10: 2138-2149.
Dell’anno F, Brunet C, Van Zyl LJ, Trindade M, Golyshin PN, Dell’anno A, Ianora A, Sansone C. 2020. Degradation of hydrocarbons and heavy metal reduction by marine bacteria in highly contaminated sediments. Microorganisms. 8: 1402.
Deng MC, Li J, Liang FR, Yi M, Xu XM, Yuan JP, Peng J, Wu CF, Wang JH. 2014. Isolation and characterization of a novel hydrocarbon-degrading bacterium Achromobacter sp. HZ01 from the crude oil-contaminated seawater at the Daya Bay, Southern China. Mar Pollut Bull. 83: 79-86.
Devereux R, Willis S. 1995. Amplification of ribosomal RNA sequences. In: Akkermans ADL, van Elsas JD, De Bruijn FJ, editors. Molecular microbial ecology manual. Vol. 3.3.1. London: Kluwer Academic Publishers. p. 1-11.
Dong C, Bai X, Sheng H, Jiao L, Zhou H, Shao Z. 2015. Distribution of PAHs and the PAH-degrading bacteria in the deep-sea sediments of the high-latitude Arctic Ocean. Biogeosciences. 12: 2163-2177.
Etkin DS. 2010. Worldwide analysis of in-port vessel operational lubricant discharges and leakages. Proceedings of 33rd AMOP Tech Semin Environ Contam Response. 1: 529-553.
Falcón LI, Noguez AM, Espinosa-Asuar L, Eguiarte LE, Souza V. 2008. Evidence of biogeography in surface ocean bacterioplankton assemblages. Mar Genom. 1: 55-61.
Gasperotti AF, Studdert CA, Revale S, Herrera Seitz MK. 2015. Draft genome sequence of Halomonas sp. KHS3, a polyaromatic hydrocarbon-chemotactic strain. Genome Announc. 3 (2): e00020-15.
Ghosal D, Ghosh S, Dutta TK, Ahn Y. 2016. Current state of knowledge in microbial degradation of polycyclic aromatic hydrocarbons (PAHs): a review. Front Microbiol. 7: 1369.
Govarthanan M, Ashraf YZ, Kamala-Kannan S, Srinivasan P, Selvankumar T, Selvam K, KIM W. 2020. Significance of allochthonous brackish water Halomonas sp. on biodegradation of low and high molecular weight polycyclic aromatic hydrocarbons. Chemosphere. 243.
Govarthanan M, Mythili R, Selvankumar T, Kamala-Kannan S, Choi D, Chang YC. 2017. Isolation and characterization of a biosurfactant-producing heavy metal resistant Rahnella sp. RM isolated from chromium-contaminated soil. Biotechnol Bioproc. 22: 186-194.
Gran-Scheuch A, Fuentes E, Bravo DM, Jiménez JC, Pérez-Donoso JM. 2017. Isolation and characterization of phenanthrene degrading bacteria from diesel fuel-contaminated Antarctic soils. Front Microbiol. 8: 1634.
Guibert LM, Loviso CL, Borglia S, Jansson JK, Dionisi HM, Lozada M. 2016. Diverse bacterial groups contribute to the alkane degradation potential of chronically polluted Subantartic coastal sediments. Microb Ecol. 71: 101-112.
Gutierrez T, Morris G, Ellis D, Mulloy B, Aitken MD. 2020. Production and characterization of a marine Halomonas surface-active exopolymer. Appl Environ Microbiol. 104: 1063-1076.
Haritash AK, Kaushik CP. 2009. Biodegradation aspects of polycyclic aromatic hydrocarbons (PAHs): a review. J Hazard Mater. 169: 1-15.
Isaac P, Bourguignon N, Maizel D, Ferrero MA. 2016. Indigenous PAH degrading bacteria in oil-polluted marine sediments from Patagonia: diversity and biotechnological properties. In: Olivera N, Libkind D, Donati E, editors. Biology and biotechnology of Patagonian microorganisms. Cham: Springer. p. 31-42.
Iyer A, Mody K, Jha B. 2006. Emulsifying properties of a marine bacterial exopolysaccharide. Enzime Microb Technol. 38: 220-222.
Izzo SA, Quintana S, Espinosa M, Babay PA, Peressutti SR. 2019. First characterization of PAH degrading bacteria from Río de la Plata and high resolution melting: an encouraging step towards bioremediation. Environ Technol. 40: 1250-1261.
Jägerbrand A, Brutemark A, Svedén JB. 2019. A review on the environmental impacts of shipping on aquatic and nearshore ecosystems. Sci Total Environ. 695: 133637.
Jain DK, Collins-Thompson DL, Lee H, Trevors JT. 1991. A drop-collapsing test for screening surfactant-producing microorganisms. J Microbiol Methods. 13: 271-279.
Johnsen AR, Wick LY, Harms H. 2005. Principles of microbial PAH-degradation in soil. Environ Pollut. 133: 71-84.
Kahng HY, Nam K, Kukor J, Yoon BJ, Lee DH, Oh DC, Kam SK, Oh KH. 2002. PAH utilization by Pseudomonas rhodesiae KK1 isolated from a former manufactured-gas plant site. Appl Microbiol Biotechnol. 60: 475-480.
Louvado A, Coelho FJRC, Gomes H, Cleary DFR, Cunha A, Gomes NCM. 2018. Independent and interactive effects of reduced seawater pH and oil contamination on subsurface sediment bacterial communities. Environ Sci Pollut Res Int. 25: 32756-32766.
Mallick S, Dutta TK. 2008. Kinetics of phenanthrene degradation by Staphylococcus sp. strain PN/Y involving 2-hydroxy-1-naphthoic acid in a novel metabolic pathway. Process Biochem. 43: 1004-1008.
McGenity TJ, Folwell BD, Mckew BA, Sanni GO. 2012. Marine crude-oil biodegradation: a central role for interspecies interactions. Aquat Biosyst. 8: 10.
Mrozik A, Piotrowska-Seget Z, Labuzek S. 2003. Bacterial degradation and bioremediation of polycyclic aromatic hydrocarbons. Pol J Environ Stud. 12: 15-25.
Muangchinda C, Chavanich S, Viyakarn V, Watanabe K, Imura S, Vangnai AS, Pinyakong O. 2015. Abundance and diversity of functional genes involved in the degradation of aromatic hydrocarbons in Antarctic soils and sediments around Syowa Station. Environ Sci Pollut Res. 22: 4725-4735.
Mulet M, David Z, Nogales B, Bosch R, Lalucat J, García-Valdés E. 2011. Pseudomonas diversity in crude-oil-contaminated intertidal sand samples obtained after the Prestige oil spill. Appl Environ Microbiol. 77: 1076-1085.
Nievas ML, Commendatore MG, Esteves JL, Bucalá V. 2005. Effect of pH modification on bilge waste biodegradation by a native microbial community. Int Biodeterior Biodegrad. 56: 151-157.
Nievas ML, Commendatore MG, Olivera NL, Esteves JL, Bucalá V. 2006. Biodegradation of bilge waste from Patagonia with an indigenous microbial community. Bioresour Technol. 97: 2280-2290.
Niosh. 1998. Polynuclear aromatic hydrocarbons by HPLC, method 5506. manual of analytical methods (NMAM). 4th ed. Issue 3, Washington: National Institute for Occupational Safety and Health (NIOSH). p. 1-9.
Olivera NL, Nievas ML, Lozada M, Del Prado G, Dionisi HM, Sineriz F. 2009. Isolation and characterization of biosurfactant-producing Alcanivorax strains: hydrocarbon accession strategies and alkane hydroxylase gene analysis. Res Microbiol. 160: 19-26.
Olivera N, Sineriz F, Breccia JD. 2005. Bacillus patagoniensis sp. nov., isolated from the rhizosphere of Atriplex lampa in Patagonia, Argentina. Int J Syst Evol Microbiol. 55: 443-447.
Pedetta A, Pouyte K, Herrera Seitz MK, Babay PA, Espinosa M, Costagliola M, Studdert CA, Peressutti SR. 2013. Phenanthrene degradation and strategies to improve its bioavailability to microorganisms isolated from brackish sediments. Int Biodeter Biodegrad. 84: 161-167.
Peng F, Liu Z, Wang L, Shao Z. 2007. An oil-degrading bacterium: Rhodococcus erythropolis strain 3C-9 and its biosurfactants. J Appl Microbiol. 102: 1603-1611.
Perelo LW. 2010. Review: In situ and bioremediation of organic pollutants in aquatic sediments. J Hazard Mater. 177: 81-89.
Peressutti SR, Alvarez HM, Pucci OH. 2003. Dynamics of Hydrocarbon-Degrading Bacteriocenosis of an Experimental Oil Pollution in Patagonian Soil. Int Biodeter Biodegrad. 52: 21-30.
Ron EZ, Rosenberg E. 2014. Enhanced bioremediation of oil spills in the sea. Curr Opin Biotechnol. 27: 191-194.
Sakshi Sing SK, Haritash AK. 2020. A comprehensive review of metabolic and genomic aspects of PAH-degradation. Arch Microbiol. 202: 2033-2058.
Scheibner MV, Herlemann DPR, Lewandowska AM, Jürgens K. 2018. Phyto- and bacterioplankton during early spring conditions in the Baltic Sea and response to short-term experimental warming. Front. Mar Sci. 5: 231.
Schlegel HG, Kaltwasser H, Gottschalk G. 1961. A submersion method for culture of hydrogen-oxidizing bacteria: growth physiological studies. Arch Microbiol. 38: 209-222.
Seo JS, Keum YS, Li QX. 2009. Bacterial Degradation of Aromatic Compounds. Int J Environ Res Public Health. 6: 278-309.
Shi K, Xue J, Xiao X, Xiao X, Qiao Y, Wu Y, Gao Y. 2019. Mechanism of degrading petroleum hydrocarbons by compound marine petroleum-degrading bacteria: surface adsorption, cell uptake, and biodegradation. Energ Fuel. 33: 11373-11379.
Silvestri GE, Berman AL. 2018. Temperatura superficial del mar en el Atlántico Sudoccidental simulada por modelos PMIP3-CMIP5: climas pasados y proyecciones futuras. X Jornadas de Ciencias del Mar. p. 275.
Song X, Xu Y, Li G, Zhang Y, Huang T, Hu Z. 2011. Isolation, characterization of Rhodococcus sp. P14 capable of degrading high-molecular-weight polycyclic aromatic hydrocarbons and aliphatic hydrocarbons. Mar Pollut Bull. 62: 2122-2128.
Tamura K, Peterson D, Peterson N, Nei M, Kumar S. 2011. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 28: 2731-2739.
Thavamani P, Megharaj M, Naidu R. 2012. Bioremediation of high molecular weight polyaromatic hydrocarbons co-contaminated with metals in liquid and soil slurries by metal tolerant PAHs degrading bacterial consortium. Biodegradation. 23: 823-835.
Tian L, Ma P, Zhong J. 2002. Kinetics and key enzyme activities of phenanthrene degradation by Pseudomonas mendocina. Process Biochem. 37: 1431-1437.
Tripathi V, Gaur VK, Dhiman N, Gautam K, Manickam N. 2020. Characterization and properties of the biosurfactant produced by PAH-degrading bacteria isolated from contaminated oily sludge environment Environ Sci Pollut Res. 27: 27268-27278.
Wilson K. 2001. Preparation of genomic DNA from bacteria. In: Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K, editors. Current protocols in molecular biology. Malden, MA: Wiley. p. 241-245.
Yuan J, Lai Q, Sun F, Zheng T, Shao Z. 2015. The diversity of PAH-degrading bacteria in a deep-sea water column above the Southwest Indian Ridge. Front Microbiol. 6: 853.
Yuan SY, Wei SH, Chang BV. 2000. Biodegradation of polycyclic aromatic hydrocarbons by a mixture culture. Chemosphere. 41: 1463-1468.
Descargas
Publicado
Número
Sección
Licencia
Derechos de autor 2020 Silvia R. Peressutti

Esta obra está bajo una licencia internacional Creative Commons Atribución-NoComercial-CompartirIgual 4.0.
Los autores de los artículos publicados en Marine and Fishery Sciences conservan los derechos de autor de sus artículos, a excepción de las imágenes de terceros y otros materiales añadidos por Marine and Fishery Sciences, que están sujetos a los derechos de autor de sus respectivos propietarios. Por lo tanto, los autores son libres de difundir y volver a publicar sus artículos, sujeto a los requisitos de los propietarios de derechos de autor de terceros y sujeto a que la publicación original sea completamente citada. Los visitantes también pueden descargar y reenviar artículos sujetos a los requisitos de citas. La capacidad de copiar, descargar, reenviar o distribuir cualquier material siempre está sujeta a los avisos de derechos de autor que se muestran. Los avisos de copyright deben mostrarse de manera prominente y no pueden borrarse, eliminarse u ocultarse, total o parcialmente. El autoalmacenamiento en servidores y repositorios de preimpresión está permitido para todas las versiones.
Esta revista ofrece a los autores una política de acceso abierto. Los usuarios pueden leer, descargar, copiar, distribuir, imprimir, buscar o vincular los textos completos de los artículos, o usarlos para cualquier otro propósito legal dentro de la licencia Creative Commons 4.0 (BY-NC-SA), sin solicitar permiso previo del editor o del autor. Esto está de acuerdo con la definición BOAI de acceso abierto.






















